Share this
Soil Microbiome Race to Measure Soil Health
by Alberto Acedo on 26/Apr/22
Enzymes in the soil may exist in a wide variety of states, like extracellular enzymes that remain active even if cells are dead.
Recently, the Soil Health Institute published an article as part of a six-part series concerning soil health assessment. This article describes the four methods currently available to commercial laboratories for measuring soil microbiome activity linked with soil health (DOI: 10.1002/crso.20099).
However, among the four methods mentioned, there are two, in particular, that stand out as the most popular: DNA-based soil measurements. Our proprietary and patented BeCrop® Technology uses this same methodology. Secondly, the potential enzyme activity measurements, which pertain to soil’s capacity to degrade specific organic molecules.
From a methodological standpoint, both tests utilize different technologies to arrive at comparable results. Let’s try to explain these differences in light of the fact that the results might appear similar, even though the reality is actually something completely opposite.
We normally understand DNA-based measurements as a way to identify microbes by sequencing, essentially reading, their DNA. This assessment can also be used to measure the soil's potential enzymatic activity by looking at the potential metabolic activities of each microbe identified, specifically as it relates to genetic coding.
In the case of the potential enzyme activity test, the technique consists of saturating a soil sample with a specific substrate and then quantifying the number of released molecules as detected by a physical-colorimetric approach. For the scientific community, these concepts are very familiar, but if you are not so scientifically inclined, perhaps a comparative example will help to clarify the differences.
Let’s think about this in terms of the automotive sector, i.e. cars. If we want to measure the transmitting power (CV joints) of an unknown car without actually seeing it, how would we accomplish this? In the case of the potential enzyme activity test, first, we would fill up the car’s gas tank, then we would start the engine and actually measure the physical noise and sound produced. Comparing a reference engine sound and the noise generated would make it possible to approximate the engine transmitting power.
By applying this same analogy to the DNA-based measurement approach, it would be similar in many ways to determining the chassis serial number or frame number. The vehicle frame number makes it possible to identify the brand and car model (much like identifying microbes in soil), and with that information, an interested party could easily reference the car guide’s database to acquire the correlating CV as described by the auto manufacturer for the specific model identified.
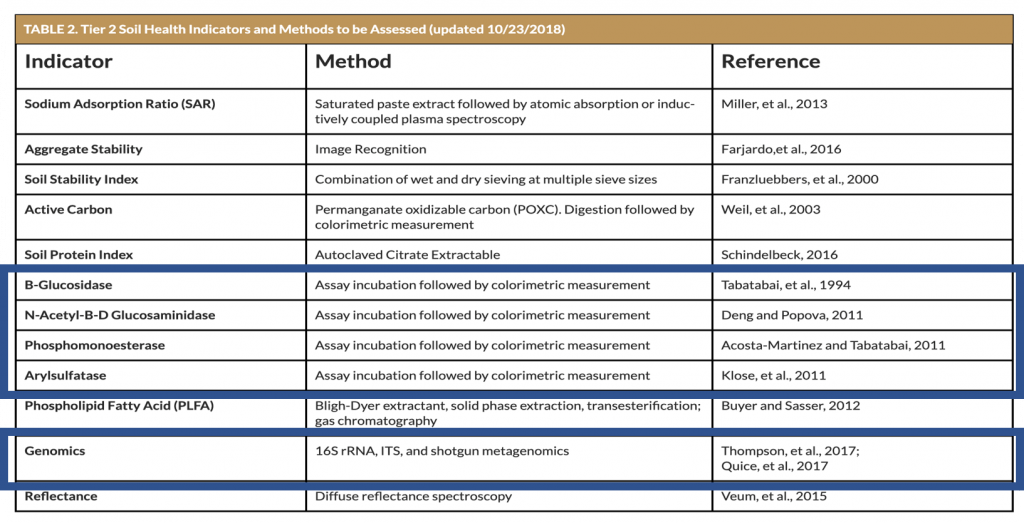
Although both approaches are totally different, the results can seem very similar. For instance, only four enzymes are studied under the potential enzyme activity approach, a relatively small number compared to the thousands of enzymes that can be identified using the DNA approach. If we go back to our automotive analogy, it can be concluded that, whereas we can easily identify the car frame number, it also now becomes possible to know all the details of the entire vehicle in question.
On top of this, enzymes in the soil may exist in a wide variety of states, like extracellular enzymes that remain active even if cells are dead. In other words, hearing the engine does not necessarily correlate to an entire, complete car. It only indicates information as it relates to the engine.
As I described in the last post (Life is DNA, and to DNA, Life Shall Return), using chemical steps in the DNA soil isolation process makes it possible to differentiate between dead and living cells. Therefore, in terms of the DNA-based approach, if we are able to determine the car frame number, it similarly indicates that the entire car exists as well.
Another interesting aspect of the enzymatic activity approach can be found in the fact that - despite it being an old technique (it is more than 100 years old) - simple interferences can dramatically affect measurement results upwards or downwards. Let’s consider, for instance, these two factors: sample conditions and storage.
Our car example will make this easier to understand. If a car is frozen, cold, or already heated up, it will not sound the same as a car under normal temperature conditions. Because DNA is a complex molecule with high chemical stability, it can survive for thousands of years. Therefore, by simply following normal sampling, storage, and shipping recommendations, deviations can easily be prevented.
Finally, the enzymatic activity approach reveals other issues related to the soil, depending on different characteristics. For example, it is commonly assumed that the pH optimum of soil enzymes used in buffers is not necessarily applicable to all soils. Just like in our car example, it’s relatively difficult to measure the transmitting power of an electric car using only sound.
Other issues that should be considered are the duration of the essays which, in turn, are linked to the complexity to replicate the essays; incomplete recovery of products; interference of DOM co-extracted, or insufficient substrate concentration that will generate different results for collected samples.
To use our car example again, it’s safe to say that all cars do not need the same amount of gas to fill up the tank. By factoring this into the equation, we can reasonably assume that an engine might sound differently if the tank is not completely full, ergo, different experiments might similarly not be comparable.
When the soil’s potential is identified through DNA sequencing in agriculture, the soil microbiome, and soil genetics, stakeholders can understand the microbial population and diversity of soil to employ the beneficial soil microorganisms to take action in agriculture.
In conclusion, there is not presently a method to directly measure the presence of enzymes in the soil. All methods measure the “potential.” But in contrast to the DNA - approach, interpretation of the potential enzyme activity approach, in relation to soil health is difficult because multiple mechanisms can drive measurement results up or down. “The lack of broad interpretability limits its use as a soil health indicator at scale,” said Elizabeth Rieke and Shannon Cappellazzi (DOI: 10.1002/crso.20099).
Originally published May 11, 2021, updated April 26, 2022.
Share this
- March 2026 (1)
- June 2025 (2)
- September 2024 (1)
- August 2024 (1)
- July 2024 (1)
- June 2024 (2)
- May 2024 (2)
- April 2024 (2)
- March 2024 (1)
- February 2024 (1)
- January 2024 (1)
- November 2023 (1)
- October 2023 (1)
- September 2023 (1)
- August 2023 (2)
- July 2023 (2)
- May 2023 (2)
- April 2023 (3)
- March 2023 (3)
- February 2023 (2)
- January 2023 (1)
- November 2022 (1)
- October 2022 (2)
- September 2022 (1)
- August 2022 (3)
- July 2022 (3)
- June 2022 (1)
- May 2022 (5)
- April 2022 (7)
- August 2021 (1)
- July 2021 (1)
- May 2021 (1)
- April 2021 (4)
- November 2020 (3)
- October 2020 (4)
- September 2020 (1)
- August 2020 (3)
- July 2020 (1)
- June 2020 (1)
- May 2020 (2)